The sodium–potassium pump (sodium–potassium adenosine triphosphatase, also known as Na+/K+-ATPase, Na+/K+ pump, or sodium–potassium ATPase) is an enzyme (an electrogenic transmembrane ATPase) found in the membrane of all animal cells. It performs several functions in cell physiology.
| Na+/K+-ATPase pump | |||||||||
|---|---|---|---|---|---|---|---|---|---|
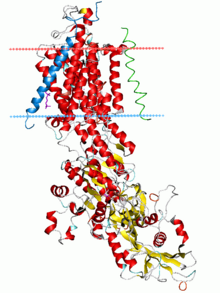 Sodium–potassium pump, E2-Pi state. Calculated hydrocarbon boundaries of the lipid bilayer are shown as blue (intracellular) and red (extracellular) planes | |||||||||
| Identifiers | |||||||||
| EC no. | 7.2.2.13 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| |||||||||


The Na+/K+-ATPase enzyme is active (i.e. it uses energy from ATP). For every ATP molecule that the pump uses, three sodium ions are exported and two potassium ions are imported.[1] Thus, there is a net export of a single positive charge per pump cycle. The net effect is an extracellular concentration of sodium ions which is 5 times the intracellular concentration, and an intracellular concentration of potassium ions which is 30 times the extracellular concentration.[1]
The sodium–potassium pump was discovered in 1957 by the Danish scientist Jens Christian Skou, who was awarded a Nobel Prize for his work in 1997. Its discovery marked an important step forward in the understanding of how ions get into and out of cells, and it has particular significance for excitable cells such as nerve cells, which depend on this pump to respond to stimuli and transmit impulses.
All mammals have four different sodium pump sub-types, or isoforms. Each has unique properties and tissue expression patterns.[2] This enzyme belongs to the family of P-type ATPases.
Function
editThe Na+/K+-ATPase helps maintain resting potential, affects transport, and regulates cellular volume.[3] It also functions as a signal transducer/integrator to regulate the MAPK pathway, reactive oxygen species (ROS), as well as intracellular calcium. In fact, all cells expend a large fraction of the ATP they produce (typically 30% and up to 70% in nerve cells) to maintain their required cytosolic Na and K concentrations.[4] For neurons, the Na+/K+-ATPase can be responsible for up to 3/4 of the cell's energy expenditure.[5] In many types of tissue, ATP consumption by the Na+/K+-ATPases have been related to glycolysis. This was first discovered in red blood cells (Schrier, 1966), but has later been evidenced in renal cells,[6] smooth muscles surrounding the blood vessels,[7] and cardiac Purkinje cells.[8] Recently, glycolysis has also been shown to be of particular importance for Na+/K+-ATPase in skeletal muscles, where inhibition of glycogen breakdown (a substrate for glycolysis) leads to reduced Na+/K+-ATPase activity and lower force production.[9][10][11]
Resting potential
editIn order to maintain the cell membrane potential, cells keep a low concentration of sodium ions and high levels of potassium ions within the cell (intracellular). The sodium–potassium pump mechanism moves 3 sodium ions out and moves 2 potassium ions in, thus, in total, removing one positive charge carrier from the intracellular space (see § Mechanism for details). In addition, there is a short-circuit channel (i.e. a highly K-permeable ion channel) for potassium in the membrane, thus the voltage across the plasma membrane is close to the Nernst potential of potassium.
Reversal potential
editEven if both K+ and Na+ ions have the same charge, they can still have very different equilibrium potentials for both outside and/or inside concentrations. The sodium-potassium pump moves toward a nonequilibrium state with the relative concentrations of Na+ and K+ for both inside and outside of cell. For instance, the concentration of K+ in cytosol is 100 mM, whereas the concentration of Na+ is 10 mM. On the other hand, in extracellular space, the usual concentration range of K+ is about 3.5-5 mM, whereas the concentration of Na+ is about 135-145 mM.[citation needed]
Transport
editExport of sodium ions from the cell provides the driving force for several secondary active transporters such as membrane transport proteins, which import glucose, amino acids and other nutrients into the cell by use of the sodium ion gradient.
Another important task of the Na+-K+ pump is to provide a Na+ gradient that is used by certain carrier processes. In the gut, for example, sodium is transported out of the reabsorbing cell on the blood (interstitial fluid) side via the Na+-K+ pump, whereas, on the reabsorbing (lumenal) side, the Na+-glucose symporter uses the created Na+ gradient as a source of energy to import both Na+ and glucose, which is far more efficient than simple diffusion. Similar processes are located in the renal tubular system.
Controlling cell volume
editFailure of the Na+-K+ pumps can result in swelling of the cell. A cell's osmolarity is the sum of the concentrations of the various ion species and many proteins and other organic compounds inside the cell. When this is higher than the osmolarity outside of the cell, water flows into the cell through osmosis. This can cause the cell to swell up and lyse. The Na+-K+ pump helps to maintain the right concentrations of ions. Furthermore, when the cell begins to swell, this automatically activates the Na+-K+ pump because it changes the internal concentrations of Na+-K+ to which the pump is sensitive.[12]
Functioning as signal transducer
editWithin the last decade[when?], many independent labs have demonstrated that, in addition to the classical ion transporting, this membrane protein can also relay extracellular ouabain-binding signalling into the cell through regulation of protein tyrosine phosphorylation. For instance, a study investigated the function of Na+/K+-ATPase in foot muscle and hepatopancreas in land snail Otala lactea by comparing the active and estivating states.[13] They concluded that reversible phosphorylation can control the same means of coordinating ATP use by this ion pump with the rates of the ATP generation by catabolic pathways in estivating O. lactea. The downstream signals through ouabain-triggered protein phosphorylation events include activation of the mitogen-activated protein kinase (MAPK) signal cascades, mitochondrial reactive oxygen species (ROS) production, as well as activation of phospholipase C (PLC) and inositol triphosphate (IP3) receptor (IP3R) in different intracellular compartments.[14]
Protein-protein interactions play a very important role in Na+-K+ pump-mediated signal transduction. For example, the Na+-K+ pump interacts directly with Src, a non-receptor tyrosine kinase, to form a signaling receptor complex.[15] Src is initially inhibited by the Na+-K+ pump. However, upon subsequent ouabain binding, the Src kinase domain is released and then activated. Based on this scenario, NaKtide, a peptide Src inhibitor derived from the Na+-K+ pump, was developed as a functional ouabain–Na+-K+ pump-mediated signal transduction.[16] Na+-K+ pump also interacts with ankyrin, IP3R, PI3K, PLCgamma1 and cofilin.[17]
Controlling neuron activity states
editThe Na+-K+ pump has been shown to control and set the intrinsic activity mode of cerebellar Purkinje neurons,[18] accessory olfactory bulb mitral cells[19] and probably other neuron types.[20] This suggests that the pump might not simply be a homeostatic, "housekeeping" molecule for ionic gradients, but could be a computation element in the cerebellum and the brain.[21] Indeed, a mutation in the Na+-K+ pump causes rapid onset dystonia-parkinsonism, which has symptoms to indicate that it is a pathology of cerebellar computation.[22] Furthermore, an ouabain block of Na+-K+ pumps in the cerebellum of a live mouse results in it displaying ataxia and dystonia.[23] Alcohol inhibits sodium–potassium pumps in the cerebellum and this is likely how it corrupts cerebellar computation and body coordination.[24][25] The distribution of the Na+-K+ pump on myelinated axons in the human brain has been demonstrated to be along the internodal axolemma, and not within the nodal axolemma as previously thought.[26] The Na+-K+ pump disfunction has been tied to various diseases, including epilepsy and brain malformations.[27]
Mechanism
editLooking at the process starting from the interior of the cell:
- The pump has a higher affinity for Na+ ions than K+ ions, thus after binding ATP, binds 3 intracellular Na+ ions.[3]
- ATP is hydrolyzed, leading to phosphorylation of the pump at a highly conserved aspartate residue and subsequent release of ADP. This process leads to a conformational change in the pump.
- The conformational change exposes the Na+ ions to the extracellular region. The phosphorylated form of the pump has a low affinity for Na+ ions, so they are released; by contrast it has high affinity for the K+ ions.
- The pump binds 2 extracellular K+ ions, which induces dephosphorylation of the pump, reverting it to its previous conformational state, thus releasing the K+ ions into the cell.
- The unphosphorylated form of the pump has a higher affinity for Na+ ions. ATP binds, and the process starts again.
Regulation
editEndogenous
editThe Na+/K+-ATPase is upregulated by cAMP.[28] Thus, substances causing an increase in cAMP upregulate the Na+/K+-ATPase. These include the ligands of the Gs-coupled GPCRs. In contrast, substances causing a decrease in cAMP downregulate the Na+/K+-ATPase. These include the ligands of the Gi-coupled GPCRs. Note: Early studies indicated the opposite effect, but these were later found to be inaccurate due to additional complicating factors. [citation needed]
The Na+/K+-ATPase is endogenously negatively regulated by the inositol pyrophosphate 5-InsP7, an intracellular signaling molecule generated by IP6K1, which relieves an autoinhibitory domain of PI3K p85α to drive endocytosis and degradation.[29]
The Na+/K+-ATPase is also regulated by reversible phosphorylation. Research has shown that in estivating animals, the Na+/K+-ATPase is in the phosphorylated and low activity form. Dephosphorylation of Na+/K+-ATPase can recover it to the high activity form.[13]
Exogenous
editThe Na+/K+-ATPase can be pharmacologically modified by administering drugs exogenously. Its expression can also be modified through hormones such as triiodothyronine, a thyroid hormone.[13][30]
For instance, Na+/K+-ATPase found in the membrane of heart cells is an important target of cardiac glycosides (for example digoxin and ouabain), inotropic drugs used to improve heart performance by increasing its force of contraction.
Muscle contraction is dependent on a 100- to 10,000-times-higher-than-resting intracellular Ca2+ concentration, which is caused by Ca2+ release from the muscle cells' sarcoplasmic reticulum. Immediately after muscle contraction, intracellular Ca2+ is quickly returned to its normal concentration by a carrier enzyme in the plasma membrane, and a calcium pump in sarcoplasmic reticulum, causing the muscle to relax.
According to the Blaustein-hypothesis,[31] this carrier enzyme (Na+/Ca2+ exchanger, NCX) uses the Na gradient generated by the Na+-K+ pump to remove Ca2+ from the intracellular space, hence slowing down the Na+-K+ pump results in a permanently elevated Ca2+ level in the muscle, which may be the mechanism of the long-term inotropic effect of cardiac glycosides such as digoxin. The problem with this hypothesis is that at pharmacological concentrations of digitalis, less than 5% of Na/K-ATPase molecules – specifically the α2 isoform in heart and arterial smooth muscle (Kd = 32 nM) – are inhibited, not enough to affect the intracellular concentration of Na+. However, apart from the population of Na/K-ATPase in the plasma membrane, responsible for ion transport, there is another population in the caveolae which acts as digitalis receptor and stimulates the EGF receptor.[32][33][34][35]
Pharmacological regulation
editIn certain conditions such as in the case of cardiac disease, the Na+/K+-ATPase may need to be inhibited via pharmacological means. A commonly used inhibitor used in the treatment of cardiac disease is digoxin (a cardiac glycoside) which essentially binds "to the extracellular part of enzyme i.e. that binds potassium, when it is in a phosphorylated state, to transfer potassium inside the cell"[36] After this essential binding occurs, a dephosphorylation of the alpha subunit occurs which reduces the effect of cardiac disease. It is via the inhibiting of the Na+/K+-ATPase that sodium levels will begin to increase within the cell which ultimately increases the concentration of intracellular calcium via the sodium-calcium exchanger. This increased presence of calcium is what allows for the force of contraction to be increased. In the case of patients where the heart is not pumping hard enough to provide what is needed for the body, use of digoxin helps to temporarily overcome this.
Discovery
editNa+/K+-ATPase was proposed by Jens Christian Skou in 1957 while working as assistant professor at the Department of Physiology, University of Aarhus, Denmark. He published his work that year.[37]
In 1997, he received one-half of the Nobel Prize in Chemistry "for the first discovery of an ion-transporting enzyme, Na+,K+-ATPase."[38]
Genes
edit- Alpha: ATP1A1, ATP1A2, ATP1A3, ATP1A4. ATP1A1 is expressed ubiquitously in vertebrates, and ATP1A3 in neural tissue. ATP1A2 is also known as "alpha(+)". ATP1A4 is specific to mammals.
- Beta: ATP1B1, ATP1B2, ATP1B3
ATP1B4, although closely related to ATP1B1, ATP1B2, and ATP1B3, lost its function as Na+/K+-ATPase beta subunit.[39]
The parallel evolution of resistance to cardiotonic steroids in many vertebrates
editSeveral studies have detailed the evolution of cardiotonic steroid resistance of the alpha-subunit gene family of Na/K-ATPase (ATP1A) in vertebrates via amino acid substitutions most often located in the first extracellular loop domain.[40][41][42][43][44][45][46] Amino acid substitutions conferring cardiotonic steroid resistance have evolved independently many times in all major groups of tetrapods.[44] ATP1A1 has been duplicated in some groups of frogs and neofunctionlised duplicates carry the same cardiotonic steroid resistance substitutions (Q111R and N122D) found in mice, rats and other muroids.[47][40][41][42]
In insects
editIn Drosophila melanogaster, the alpha-subunit of Na+/K+-ATPase has two paralogs, ATPα (ATPα1) and JYalpha (ATPα2), resulting from an ancient duplication in insects.[48] In Drosophila, ATPα1 is ubiquitously and highly expressed, whereas ATPα2 is most highly expressed in male testes and is essential for male fertility. Insects have at least one copy of both genes, and occasionally duplications. Low expression of ATPα2 has also been noted in other insects. Duplications and neofunctionalization of ATPα1 have been observed in insects that are adapted to cardiotonic steroid toxins such as cardenolides and bufadienolides.[48][49][50] Insects adapted to cardiotonic steroids typically have a number of amino acid substitutions, most often in the first extra-cellular loop of ATPα1, that confer resistance to cardiotonic steroid inhibition.[51][52]
See also
editReferences
edit- ^ a b Gagnon KB, Delpire E (2021). "Sodium Transporters in Human Health and Disease (Figure 2)". Frontiers in Physiology. 11: 588664. doi:10.3389/fphys.2020.588664. PMC 7947867. PMID 33716756.
- ^ Clausen MV, Hilbers F, Poulsen H (June 2017). "The Structure and Function of the Na,K-ATPase Isoforms in Health and Disease". Frontiers in Physiology. 8: 371. doi:10.3389/fphys.2017.00371. PMC 5459889. PMID 28634454.
- ^ a b Hall JE, Guyton AC (2006). Textbook of medical physiology. St. Louis, Mo: Elsevier Saunders. ISBN 978-0-7216-0240-0.
- ^ Voet D, Voet JG (December 2010). "Section 20-3: ATP-Driven Active Transport". Biochemistry (4th ed.). John Wiley & Sons. p. 759. ISBN 978-0-470-57095-1.
- ^ Howarth C, Gleeson P, Attwell D (July 2012). "Updated energy budgets for neural computation in the neocortex and cerebellum". Journal of Cerebral Blood Flow and Metabolism. 32 (7): 1222–32. doi:10.1038/jcbfm.2012.35. PMC 3390818. PMID 22434069.
- ^ Sanders MJ, Simon LM, Misfeldt DS (March 1983). "Transepithelial transport in cell culture: bioenergetics of Na-, D-glucose-coupled transport". Journal of Cellular Physiology. 114 (3): 263–6. doi:10.1002/jcp.1041140303. PMID 6833401. S2CID 22543559.
- ^ Lynch RM, Paul RJ (March 1987). "Compartmentation of carbohydrate metabolism in vascular smooth muscle". The American Journal of Physiology. 252 (3 Pt 1): C328-34. doi:10.1152/ajpcell.1987.252.3.c328. PMID 3030131.
- ^ Glitsch HG, Tappe A (January 1993). "The Na+/K+ pump of cardiac Purkinje cells is preferentially fuelled by glycolytic ATP production". Pflügers Archiv. 422 (4): 380–5. doi:10.1007/bf00374294. PMID 8382364. S2CID 25076348.
- ^ Dutka TL, Lamb GD (September 2007). "Na+-K+ pumps in the transverse tubular system of skeletal muscle fibers preferentially use ATP from glycolysis". American Journal of Physiology. Cell Physiology. 293 (3): C967-77. doi:10.1152/ajpcell.00132.2007. PMID 17553934. S2CID 2291836.
- ^ Watanabe D, Wada M (December 2019). "Effects of reduced muscle glycogen on excitation-contraction coupling in rat fast-twitch muscle: a glycogen removal study". Journal of Muscle Research and Cell Motility. 40 (3–4): 353–364. doi:10.1007/s10974-019-09524-y. PMID 31236763. S2CID 195329741.
- ^ Jensen R, Nielsen J, Ørtenblad N (February 2020). "Inhibition of glycogenolysis prolongs action potential repriming period and impairs muscle function in rat skeletal muscle". The Journal of Physiology. 598 (4): 789–803. doi:10.1113/JP278543. PMID 31823376. S2CID 209317559.
- ^ Armstrong CM (May 2003). "The Na/K pump, Cl ion, and osmotic stabilization of cells". Proceedings of the National Academy of Sciences of the United States of America. 100 (10): 6257–62. Bibcode:2003PNAS..100.6257A. doi:10.1073/pnas.0931278100. PMC 156359. PMID 12730376.
- ^ a b c Ramnanan CJ, Storey KB (February 2006). "Suppression of Na+/K+-ATPase activity during estivation in the land snail Otala lactea". The Journal of Experimental Biology. 209 (Pt 4): 677–88. doi:10.1242/jeb.02052. PMID 16449562. S2CID 39271006.
- ^ Yuan Z, Cai T, Tian J, Ivanov AV, Giovannucci DR, Xie Z (September 2005). "Na/K-ATPase tethers phospholipase C and IP3 receptor into a calcium-regulatory complex". Molecular Biology of the Cell. 16 (9): 4034–45. doi:10.1091/mbc.E05-04-0295. PMC 1196317. PMID 15975899.
- ^ Tian J, Cai T, Yuan Z, Wang H, Liu L, Haas M, et al. (January 2006). "Binding of Src to Na+/K+-ATPase forms a functional signaling complex". Molecular Biology of the Cell. 17 (1): 317–26. doi:10.1091/mbc.E05-08-0735. PMC 1345669. PMID 16267270.
- ^ Li Z, Cai T, Tian J, Xie JX, Zhao X, Liu L, et al. (July 2009). "NaKtide, a Na/K-ATPase-derived peptide Src inhibitor, antagonizes ouabain-activated signal transduction in cultured cells". The Journal of Biological Chemistry. 284 (31): 21066–76. doi:10.1074/jbc.M109.013821. PMC 2742871. PMID 19506077.
- ^ Lee K, Jung J, Kim M, Guidotti G (January 2001). "Interaction of the alpha subunit of Na,K-ATPase with cofilin". The Biochemical Journal. 353 (Pt 2): 377–85. doi:10.1042/0264-6021:3530377. PMC 1221581. PMID 11139403.
- ^ Forrest MD, Wall MJ, Press DA, Feng J (December 2012). "The sodium-potassium pump controls the intrinsic firing of the cerebellar Purkinje neuron". PLOS ONE. 7 (12): e51169. Bibcode:2012PLoSO...751169F. doi:10.1371/journal.pone.0051169. PMC 3527461. PMID 23284664.
- ^ Zylbertal A, Kahan A, Ben-Shaul Y, Yarom Y, Wagner S (December 2015). "Prolonged Intracellular Na+ Dynamics Govern Electrical Activity in Accessory Olfactory Bulb Mitral Cells". PLOS Biology. 13 (12): e1002319. doi:10.1371/journal.pbio.1002319. PMC 4684409. PMID 26674618.
- ^ Zylbertal A, Yarom Y, Wagner S (2017). "The Slow Dynamics of Intracellular Sodium Concentration Increase the Time Window of Neuronal Integration: A Simulation Study". Frontiers in Computational Neuroscience. 11: 85. doi:10.3389/fncom.2017.00085. PMC 5609115. PMID 28970791.
- ^ Forrest MD (December 2014). "The sodium-potassium pump is an information processing element in brain computation". Frontiers in Physiology. 5 (472): 472. doi:10.3389/fphys.2014.00472. PMC 4274886. PMID 25566080.
- ^ Cannon SC (July 2004). "Paying the price at the pump: dystonia from mutations in a Na+/K+-ATPase". Neuron. 43 (2): 153–4. doi:10.1016/j.neuron.2004.07.002. PMID 15260948.
- ^ Calderon DP, Fremont R, Kraenzlin F, Khodakhah K (March 2011). "The neural substrates of rapid-onset Dystonia-Parkinsonism". Nature Neuroscience. 14 (3): 357–65. doi:10.1038/nn.2753. PMC 3430603. PMID 21297628.
- ^ Forrest MD (April 2015). "Simulation of alcohol action upon a detailed Purkinje neuron model and a simpler surrogate model that runs >400 times faster". BMC Neuroscience. 16 (27): 27. doi:10.1186/s12868-015-0162-6. PMC 4417229. PMID 25928094.
- ^ Forrest M (4 April 2015). "The Neuroscience Reason We Fall Over When Drunk". Science 2.0. Retrieved 30 May 2018.
- ^ Young EA, Fowler CD, Kidd GJ, Chang A, Rudick R, Fisher E, Trapp BD (April 2008). "Imaging correlates of decreased axonal Na+/K+ ATPase in chronic multiple sclerosis lesions". Annals of Neurology. 63 (4): 428–35. doi:10.1002/ana.21381. PMID 18438950. S2CID 14658965.
- ^ Smith RS, Florio M, Akula SK, Neil JE, Wang Y, Hill RS, et al. (June 2021). "Early role for a Na+,K+-ATPase (ATP1A3) in brain development". Proceedings of the National Academy of Sciences of the United States of America. 118 (25): e2023333118. Bibcode:2021PNAS..11823333S. doi:10.1073/pnas.2023333118. PMC 8237684. PMID 34161264.
- ^ Burnier M (2008). Sodium In Health And Disease. CRC Press. p. 15. ISBN 978-0-8493-3978-3.
- ^ Chin AC, Gao Z, Riley AM, Furkert D, Wittwer C, Dutta A, et al. (October 2020). "The inositol pyrophosphate 5-InsP7 drives sodium-potassium pump degradation by relieving an autoinhibitory domain of PI3K p85α". Science Advances. 6 (44): eabb8542. Bibcode:2020SciA....6.8542C. doi:10.1126/sciadv.abb8542. PMC 7608788. PMID 33115740. S2CID 226036261.
- ^ Lin HH, Tang MJ (January 1997). "Thyroid hormone upregulates Na,K-ATPase α and β mRNA in primary cultures of proximal tubule cells". Life Sciences. 60 (6): 375–382. doi:10.1016/S0024-3205(96)00661-3. PMID 9031683.
- ^ Blaustein MP (May 1977). "Sodium ions, calcium ions, blood pressure regulation, and hypertension: a reassessment and a hypothesis". The American Journal of Physiology. 232 (5): C165-73. doi:10.1152/ajpcell.1977.232.5.C165. PMID 324293. S2CID 9814212.
- ^ Schoner W, Scheiner-Bobis G (September 2008). "Role of endogenous cardiotonic steroids in sodium homeostasis". Nephrology, Dialysis, Transplantation. 23 (9): 2723–9. doi:10.1093/ndt/gfn325. PMID 18556748.
- ^ Blaustein MP, Hamlyn JM (December 2010). "Signaling mechanisms that link salt retention to hypertension: endogenous ouabain, the Na+ pump, the Na+/Ca2+ exchanger and TRPC proteins". Biochimica et Biophysica Acta (BBA) - Molecular Basis of Disease. 1802 (12): 1219–29. doi:10.1016/j.bbadis.2010.02.011. PMC 2909369. PMID 20211726.
- ^ Fuerstenwerth H (2014). "On the differences between ouabain and digitalis glycosides". American Journal of Therapeutics. 21 (1): 35–42. doi:10.1097/MJT.0b013e318217a609. PMID 21642827. S2CID 20180376.
- ^ Pavlovic D (2014). "The role of cardiotonic steroids in the pathogenesis of cardiomyopathy in chronic kidney disease". Nephron Clinical Practice. 128 (1–2): 11–21. doi:10.1159/000363301. PMID 25341357. S2CID 2066801.
- ^ "Na+/K+-ATPase and inhibitors (Digoxin)". Pharmacorama. Archived from the original on 2020-09-28. Retrieved 2019-11-08.
- ^ Skou JC (February 1957). "The influence of some cations on an adenosine triphosphatase from peripheral nerves". Biochimica et Biophysica Acta. 23 (2): 394–401. doi:10.1016/0006-3002(57)90343-8. PMID 13412736. S2CID 32516710.
- ^ "The Nobel Prize in Chemistry 1997". NobelPrize.org. Nobel Media AB. 15 October 1997.
- ^ "ATPase Na+/K+ transporting subunits (ATP1)". HGNC. Retrieved 26 June 2024.
- ^ a b Moore, David J.; Halliday, Damien C. T.; Rowell, David M.; Robinson, Anthony J.; Keogh, J. Scott (2009-08-23). "Positive Darwinian selection results in resistance to cardioactive toxins in true toads (Anura: Bufonidae)". Biology Letters. 5 (4): 513–516. doi:10.1098/rsbl.2009.0281. ISSN 1744-9561. PMC 2781935. PMID 19465576.
- ^ a b Hernández Poveda M (2022) Convergent evolution of neo-functionalized duplications of ATP1A1 in dendrobatid and grass frogs. MS Thesis Dissertation. Universidad de los Andes
- ^ a b Mohammadi, Shabnam; Yang, Lu; Harpak, Arbel; Herrera-Álvarez, Santiago; Rodríguez-Ordoñez, María del Pilar; Peng, Julie; Zhang, Karen; Storz, Jay F.; Dobler, Susanne; Crawford, Andrew J.; Andolfatto, Peter (2021-06-21). "Concerted evolution reveals co-adapted amino acid substitutions in frogs that prey on toxic toads". Current Biology. 31 (12): 2530–2538.e10. doi:10.1016/j.cub.2021.03.089. ISSN 0960-9822. PMC 8281379. PMID 33887183.
- ^ Mohammadi, Shabnam; Brodie, Edmund D.; Neuman-Lee, Lorin A.; Savitzky, Alan H. (2016-05-01). "Mutations to the cardiotonic steroid binding site of Na+/K+-ATPase are associated with high level of resistance to gamabufotalin in a natricine snake". Toxicon. 114: 13–15. doi:10.1016/j.toxicon.2016.02.019. ISSN 0041-0101. PMID 26905927.
- ^ a b Mohammadi, Shabnam; Herrera-Álvarez, Santiago; Yang, Lu; Rodríguez-Ordoñez, María del Pilar; Zhang, Karen; Storz, Jay F.; Dobler, Susanne; Crawford, Andrew J.; Andolfatto, Peter (2022-08-16). "Constraints on the evolution of toxin-resistant Na,K-ATPases have limited dependence on sequence divergence". PLOS Genetics. 18 (8): e1010323. doi:10.1371/journal.pgen.1010323. ISSN 1553-7390. PMC 9462791. PMID 35972957.
- ^ Mohammadi, Shabnam; Özdemir, Halil İbrahim; Ozbek, Pemra; Sumbul, Fidan; Stiller, Josefin; Deng, Yuan; Crawford, Andrew J; Rowland, Hannah M; Storz, Jay F; Andolfatto, Peter; Dobler, Susanne (2022-12-06). "Epistatic Effects Between Amino Acid Insertions and Substitutions Mediate Toxin resistance of Vertebrate Na+,K+-ATPases". Molecular Biology and Evolution. 39 (12): msac258. doi:10.1093/molbev/msac258. ISSN 0737-4038. PMC 9778839. PMID 36472530.
- ^ Ujvari, Beata; Mun, Hee-chang; Conigrave, Arthur D.; Bray, Alessandra; Osterkamp, Jens; Halling, Petter; Madsen, Thomas (January 2013). "Isolation Breeds Naivety: Island Living Robs Australian Varanid Lizards of Toad-Toxin Immunity Via Four-Base-Pair Mutation". Evolution. 67 (1): 289–294. doi:10.1111/j.1558-5646.2012.01751.x. PMID 23289579.
- ^ Price, Elmer M.; Lingrel, Jerry B. (1988-11-01). "Structure-function relationships in the sodium-potassium ATPase .alpha. subunit: site-directed mutagenesis of glutamine-111 to arginine and asparagine-122 to aspartic acid generates a ouabain-resistant enzyme". Biochemistry. 27 (22): 8400–8408. doi:10.1021/bi00422a016. ISSN 0006-2960. PMID 2853965.
- ^ a b Zhen, Ying; Aardema, Matthew L.; Medina, Edgar M.; Schumer, Molly; Andolfatto, Peter (2012-09-28). "Parallel Molecular Evolution in an Herbivore Community". Science. 337 (6102): 1634–1637. Bibcode:2012Sci...337.1634Z. doi:10.1126/science.1226630. ISSN 0036-8075. PMC 3770729. PMID 23019645.
- ^ Yang, L.; Ravikanthachari, N.; Mariño-Pérez, R.; Deshmukh, R.; Wu, M.; Rosenstein, A.; Kunte, K.; Song, H.; Andolfatto, P. (2019). "Predictability in the evolution of Orthopteran cardenolide insensitivity". Philosophical Transactions of the Royal Society of London, Series B. 374 (1777): 20180246. doi:10.1098/rstb.2018.0246. PMC 6560278. PMID 31154978.
- ^ Petschenka Georg, Vera Wagschal, Michael von Tschirnhaus, Alexander Donath, Susanne Dobler 2017 Petschenka, G.; Wagschal, V.; von Tschirnhaus, M.; Donath, A.; Dobler, S. (2017). "Convergently Evolved Toxic Secondary Metabolites in Plants Drive the Parallel Molecular Evolution of Insect Resistance". The American Naturalist. 190 (S1): S29–S43. doi:10.1086/691711. PMID 28731826. S2CID 3908073.
- ^ Labeyrie E, Dobler S (2004). "Molecular adaptation of Chrysochus leaf beetles to toxic compounds in their food plants". Molecular Biology and Evolution. 21 (2): 218–21. doi:10.1093/molbev/msg240. PMID 12949136.
- ^ Dobler, Susanne; Dalla, Safaa; Wagschal, Vera; Agrawal, Anurag A. (2012). "Community-wide convergent evolution in insect adaptation to toxic cardenolides by substitutions in the Na,K-ATPase". Proceedings of the National Academy of Sciences. 109 (32): 13040–13045. doi:10.1073/pnas.1202111109. PMC 3420205. PMID 22826239.
External links
edit- Sodium,+Potassium+ATPase at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- RCSB Protein Data Bank: Sodium–Potassium Pump
- A video by Khan Academy.